Chromatin Labeling and Imaging
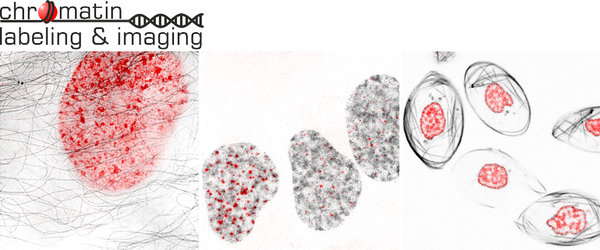
Localization of chromosomes and even specific DNA sequences in the nuclear space play crucial role in the gene expression regulation during cell cycle, oncogenesis and neuronal signalling. Nucleus morphology has been used for diagnostics of various types of cancers for decades. However, these experiments are still performed mostly on fixed cells and suffer from such artifacts as structure modification or loss during sample preparation. The modern super-resolution fluorescent microscopy techniques allow imaging of nucleus in living cells at resolution which is only slightly lower compared to electron microscopy (Figure 1), thus providing an obvious advantage. For example, by using quantitative super-resolution microscopy it has been recently demonstrated that chromatin packing density (number and size of nanodomains) strongly correlates with the pluripotency potential of induced pluripotent stem cells.


Figure 2. Example image of a DNA binding probe in the nucleus of a living cell showing more detailed structure in the STED image as compared to the confocal. Scale bar = 2 µm.
Fluorophores for super-resolution microscopy
A wider application of super-resolution imaging requires development of fluorescent probes for specific labelling. These probes are constructed by coupling fluorescent dye to a small molecule ligand, targeting nuclear components of interest. A large variety of inhibitors, cofactor analogues and binders targeting proteins and DNA are available, thus successful development of chromatin probes is feasible. However direct tagging of chromatin proteins, specific DNA sequences or DNA modifications impose a challenge of identifying correct combination of targeting moiety, linker and fluorophore.

Major interests
The laboratory combines chemistry, biology and microscopy techniques to reach the following aims:
- Create of the new biocompatible fluorescent probes targeting biomolecules.
- Explore new labeling methods for efficient labeling of biomolecules.
- Apply of the newly developed probes for imaging and establishment of image analysis pipelines.
- Image of dynamic processes in living cells during cell cycle or external stimulation.



