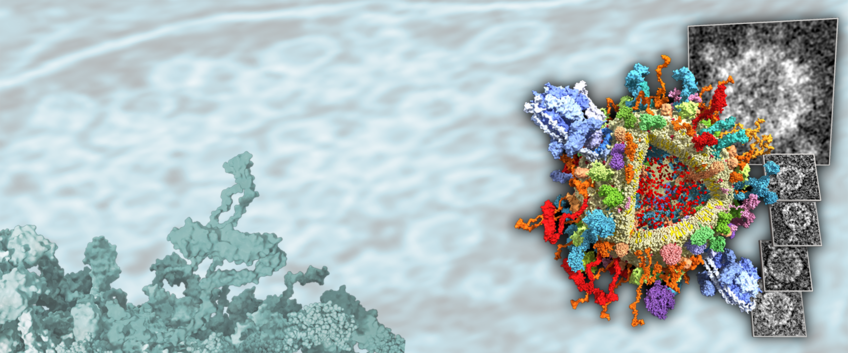
Theoretical and Computational Biophysics
Our research aims at an understanding of the physics and function of proteins, protein complexes, and other biomolecular structures at the atomic level. For this purpose, complex computer simulations of the atomistic dynamics are carried out. Read more about our research projects at our RESEARCH website.
Open Positions
We are always looking for highly motivated scientists, possessing a strong background in physics, physical chemistry, mathematics and/or biomolecular simulations. If you are interested in driving research projects forward which are conducted at our department please submit your application documents (including motivation letter, CV, publication list, certificates) preferrably as single PDF file via email to office.theor_comp_biophys@mpinat.mpg.de or follow the instructions in the job announcements.
... in the field of
Theory and Methods for Non-equilibrium Atomistic Simulations of Complex Biomolecules in the research team of Helmut Grubmüller
more
Our Research Groups
Group leader: Helmut Grubmüller
Group leader: Bert de Groot
Group leader: Aljaz Godec
Helmut's scientific writing guidelines
Struggling with writing your paper draft or thesis? Here’s advice my students found helpful over the years. It’s version 1.0 -- Comments, suggestions, corrections highly appreciated!
more
Press releases & research news
Research Group of Helmut Grubmüller
The German National Academy of Sciences Leopoldina honors the biophysicist for his outstanding scientific achievements and the special expertise in his field of research. (in German)
more
Show more
Research Group of Bert de Groot
In Parkinson’s patients, alpha-synuclein proteins clump together to form fibrils, which presumably damage nerve cells. A research team has now shown how lipids bind to these fibrils and influence their arrangement. They also demonstrated how the drug candidate anle138b attaches to the lipidic fibrils. The findings could open up new diagnostic and therapeutic approaches.
more
Show more
Research Group of Aljaz Godec
If you take a coin out of an ice bath, it warms up over time. Likewise, a hot coin that you have just taken out of a sauna cools down. The fact that systems, here the coin, thermally adapt to their environment is due to the heat flow that results from temperature differences.
more
Show more