Electron-Spin Resonance Spectroscopy
The research group is dedicated to modern electron-spin resonance (ESR/EPR) spectroscopy, from high-frequency EPR to methods at the interface with nuclear magnetic resonance (NMR) and their applications in biological science. We have been investigating how to excite and detect paramagnetic centers and their coupled nuclear spins with microwave (mw) and radio frequency (rf) pulses to achieve information on active sites of proteins or on the global structure of biomolecules. One main strategy is to transfer the much larger polarization of electron spins to nuclear spins. To this end, developments of pulse schemes at high magnetic fields (≥ 3 Tesla) and corresponding resonant mw frequencies are in focus. The most representative applications are the investigation of enzymatic reactions involving paramagnetic intermediates, particularly the proton-coupled electron transfer (PCET) in E. coli ribonucleotide reductase (RNR), and the exploration of long-range structural information in nucleic acids and transmembrane peptides by pulsed dipolar spectroscopy.
ENDOR for information from the atomic to nano scale
Electron-nuclear double resonance (ENDOR), particularly at high frequencies, permits to determine the distance as well as the orientation of nuclear spins in the ligand sphere (r < 1.5 nm) of a paramagnetic centre. For instance, hydrogen bond interactions in the active states of an enzyme can be detected that are not easily accessible by other techniques such as X-ray, cryo-EM or NMR. The combination of ENDOR with quantum chemical calculations becomes a unique tool to elucidate the structure of complex intermediates during enzymatic catalysis. We have recently demonstrated the spectroscopic detection of hydrogen bond networks in proteins via 2H ENDOR. The 17O detection of water molecules participating in proton-coupled electron transfer (PCET).

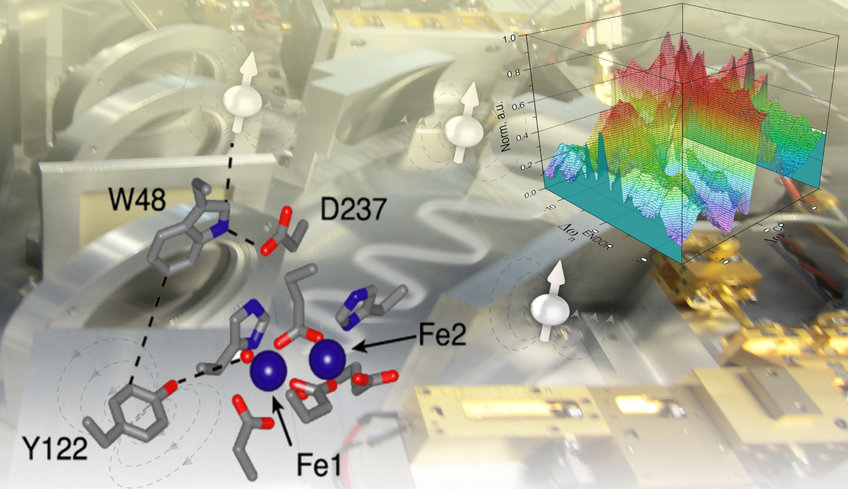
Representative applications of ENDOR spectroscopy for studiesof the PCET in E. coli RNR. Top (right): Docking model of subunits a2 (red) and b2 (blue) based on PDB structures 4r1r and 1mxr. Arrow indicates the proposed PCET pathway within one ab pair, involving four essential tyrosines, Y730/Y731 in a and Y356/Y122 in b. Top: (left) 2H ENDOR experiments (Argirevic et al, JACS 2012, Nick et al. JACS 2015) inferred a H bond network between the intermediates in a. Bottom: (right) 263 GHz 1H ENDOR spectrum of Y122• for excitation at B||gz and simulations with hf tensors of contributing nuclei (colors) (Tkach et al, J. Magn. Reson. 2019). Bottom (left): Detection of water molecules H bonded to the three Y intermediates by 17O ENDOR (Hecker et al, JACS 2021).
19F ENDOR for distance measurements: Fluorine labelling has been established in NMR as powerful and selective tool to extract structural information in biological systems. 19F can also be detected in ENDOR by taking advantage of the large electron spin polarization. We have recently shown that 19F ENDOR spectroscopy can be used to complement EPR distance measurements in the angstrom to nanometer range by introducing a nitroxide and a 19F in the molecule of study. This can be performed in nucleic acids as well as in enzyme and proteins. New labelling procedures to site-selectively introduce unnatural amino acids in proteins open the route for this type of studies. Currently, we are employing this method to address the open question of the PCET mechanism in RNR across the subunit interface. Measurement of the distance across the subunit interface in active E.coli RNR will provide important new structural information.

Representative application of 19F ENDOR to measure inter spin distances between a TEMPO label and 19F in two RNA duplexes. The investigated distance range using Mims ENDOR (pulse sequence in the figure) is displayed in the plot. Below about 6 Å, the hyperfine interaction is well detectable but the point-dipole model breaks down. The upper distance limit is dictated by the resolution of the ENDOR experiment. Figure adapted from A. Meyer et al, Angew. Chemie 2020.
Development of ENDOR pulse sequences (DFG SPP-1601): In the past year, we have started a detailed investigation of ENDOR pulse sequences to optimize their performance for different interaction ranges and different nuclear spins. We are examining the effect of nuclear spin relaxation and proposing new sequences based on electron spin lock and electron-nuclear cross polarization, to circumvent signal saturation effects. We have been implementing analytical and numerical tools to predict ENDOR signals based on a static spin Hamiltonian as well as density matrix formalism, product and single transition operators (Bejenke et al, Mol. Physics 2020). The goal is to predict the efficiency of more complex pulse sequences in powder patterns under real sample conditions, i.e. low temperatures and high electron spin polarization.
Statistical methods for ENDOR (DFG-CRC 1456): Mathematical methods are gaining increasing importance in various fields of spectroscopy to extract weak signal from noise and baseline distortions. This applies also for ENDOR spectra, where resonances are often very broad and close to the baseline. In order to address this issue, we have started a collaboration with the group of S. Huckemann (Dept. Mathematical Stochastics, Göttingen) and Yvo Pokern (UCL London) focusing on statistical analysis of ENDOR data. This approach allows for the separation of signal drifts during long acquisition times and for the determination of the most-likely ENDOR signal with uncertainties, paving the way for more quantitative data analysis (Pokern et al., PNAS DOI 10.1073/pnas.2023615118).
Dynamic Nuclear Polarization (DNP)
Adressing the sensitivity issue of nuclear magnetic resonance has been a long-standing goal and has continuously driven the community towards higher static magnetic fields, new detection technologies, and more complex radio frequency excitation schemes. Dynamic nuclear polarization (DNP) can significantly enhance the NMR signal by transferring the higher polarization from a polarizing agent (namely a paramagnetic center) to the target nuclei. In the liquid state at room temperature, the addition of stable organic radicals and irradiation of their electronic transitions via microwave can lead to NMR-signal enhancements up to two or three orders of magnitude. Our goal is to integrate DNP for routine NMR spectroscopy in the liquid state at room temperature, and then contribute to the development of the next generation of NMR methods.
The polarization transfer process critically depends on the chosen polarizing agent / target system as well as on the external magnetic field. Specifically, molecular diffusion, molecular collisions, structural reorientations, and transient bonding/complexations, all contribute to the DNP efficiency by modulating the hyperfine coupling between electron and nuclei. We aim at a deep understanding of the polarization transfer mechanisms in the liquid state. By using DNP instruments at different magnetic fields (0.34 T, 1.2 T, 3.4 T, and 9.4 T) we are characterizing new targets and polarizing agents to identify suited conditions to boost the NMR enhancements.

(a) Schematic of a DNP experiment and its mechanism in liquids. (b) Measured 13C DNP enhancement of the model system CCl4, reaching a factor of about 103 (Liu et al, Nature Chemistry 2017). Right: 13C NMR spectra obtained at 9.4 Tesla and room temperature in small organic molecules under DNP condition (from Orlando et al, Angew. Chemie 2019).
Press releases & research news



